SummaryRMgm-338
|
||||||||
 *RMgm-338
*RMgm-338| Successful modification | The parasite was generated by the genetic modification |
| The mutant contains the following genetic modification(s) | Gene tagging |
| Reference (PubMed-PMID number) |
Reference 1 (PMID number) : 20478340 |
| MR4 number | |
| top of page | |
| Parent parasite used to introduce the genetic modification | |
| Rodent Malaria Parasite | P. berghei |
| Parent strain/line | P. berghei ANKA |
| Name parent line/clone | P. berghei ANKA 2.34 |
| Other information parent line | P. berghei ANKA 2.34 is a cloned, gametocyte producer line of the ANKA strain (PubMed: PMID: 15137943). |
| top of page | |
| The mutant parasite was generated by | |
| Name PI/Researcher | S. Déchamps; K. Wengelnik; H.J. Vial; L. Gannoun-Zaki |
| Name Group/Department | Dynamique des Interactions Membranaires Normales et Pathologiques |
| Name Institute | Université Montpellier |
| City | Montpellier |
| Country | France |
| top of page | |
| Name of the mutant parasite | |
| RMgm number | RMgm-338 |
| Principal name | CK-GFP |
| Alternative name | |
| Standardized name | |
| Is the mutant parasite cloned after genetic modification | No |
| top of page | |
| Phenotype | |
| Asexual blood stage | Western analysis using GFP-antibodies and fluorescence microscopy showed CK-GFP expression in (the cytoplasm of) blood stages. |
| Gametocyte/Gamete | Not tested |
| Fertilization and ookinete | Not tested |
| Oocyst | Not tested |
| Sporozoite | Not tested |
| Liver stage | Not tested |
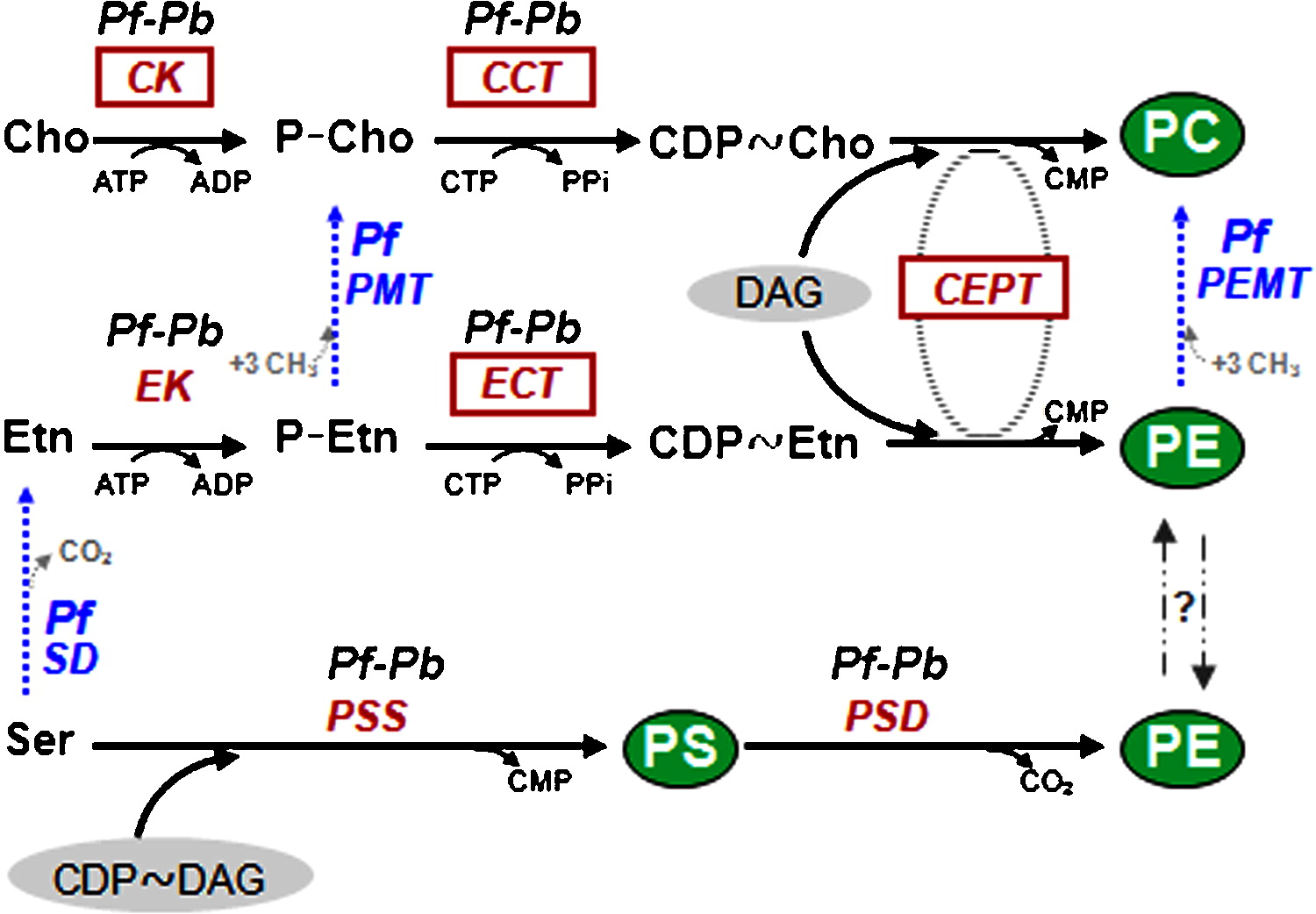
| Additional remarks phenotype | Mutant/mutation In P. berghei CK, the Brenner's phosphotransferase motif FCHNDLQENN is found. The choline kinase motif hhDhExxxxNxxxxDhxNhhxE and the conserved hydrophobic residues (Trp394, Trp397, and Tyr416) are present and make it likely that the ck gene encodes a choline kinase enzyme. An additional route termed serine decarboxylation-phosphoethanolamine methylation (SDPM) pathway has been identified in P. falciparum. In this plant-like pathway that connects different routes, the polar head groups of PE and PC are synthesized from serine that is either directly imported from the host or obtained through degradation of host cell haemoglobin. However, phosphoethanolamine N80 methyltransferase (PMT) activity is absent from P. berghei and an ortholog to the P. falciparum pmt gene (MAL13P1.214) could not be identified in the genome of P. berghei and other rodent malaria parasites.
FIG 1. Biosynthesis pathways for PC, PE and PS in Plasmodium. Other mutants |
 Tagged: Mutant parasite with a tagged gene
Tagged: Mutant parasite with a tagged gene| top of page | |||||||||||||||||||||||||||
| Details of the target gene | |||||||||||||||||||||||||||
| Gene Model of Rodent Parasite | PBANKA_1040100 | ||||||||||||||||||||||||||
| Gene Model P. falciparum ortholog | PF3D7_1401800 | ||||||||||||||||||||||||||
| Gene product | choline kinase | ||||||||||||||||||||||||||
| Gene product: Alternative name | CK | ||||||||||||||||||||||||||
| top of page | |||||||||||||||||||||||||||
| Details of the genetic modification | |||||||||||||||||||||||||||
| Name of the tag | GFP-mutant3 | ||||||||||||||||||||||||||
| Details of tagging | C-terminal | ||||||||||||||||||||||||||
| Additional remarks: tagging | |||||||||||||||||||||||||||
| Commercial source of tag-antibodies | |||||||||||||||||||||||||||
| Type of plasmid/construct | Plasmid single cross-over | ||||||||||||||||||||||||||
| PlasmoGEM (Sanger) construct/vector used | No | ||||||||||||||||||||||||||
| Modified PlasmoGEM construct/vector used | No | ||||||||||||||||||||||||||
| Plasmid/construct map | |||||||||||||||||||||||||||
| Plasmid/construct sequence | |||||||||||||||||||||||||||
| Restriction sites to linearize plasmid | EcoRV | ||||||||||||||||||||||||||
| Selectable marker used to select the mutant parasite | tgdhfr | ||||||||||||||||||||||||||
| Promoter of the selectable marker | pbdhfr | ||||||||||||||||||||||||||
| Selection (positive) procedure | pyrimethamine | ||||||||||||||||||||||||||
| Selection (negative) procedure | No | ||||||||||||||||||||||||||
| Additional remarks genetic modification | The single cross-over event result in replacement of the endogenous gene with a ck gene fused to a GFP-tag at the C-terminus. To generate the pDR-CK-GFP construct, the nucleotide sequence was modified without changing the amino acid sequence to add an EcoRV site and amplified using genomic DNA as a template with the following primer pairs: p5/p6, and p7/p8. Amplification using primers p5 and p8 gave the final fragment of 1110 bp (nucleotides 214-1323 of the Pbck ORF) that was inserted in pDR009. | ||||||||||||||||||||||||||
| Additional remarks selection procedure | |||||||||||||||||||||||||||
| |||||||||||||||||||||||||||
| top of page | |||||||||||||||||||||||||||